Disentangling the roles of TM8-like genes in reproductive organ development in Gnetum montanum Markgr.
-
Abstract:
TM8 genes, belonging to the ancient subfamily of type Ⅱ MADS-box genes, have been lost in various angiosperm lineages but have undergone a dramatic expansion in gymnosperms. While TM8 genes are known to participate in female flower development in angiosperms, their roles in gymnosperms remain poorly understood. In this study, three TM8-like genes, including one gene with two transcripts, were characterized in Gnetum montanum Markgr. using fluorescence in situ hybridization (FISH) and transgenic experiments. Results indicated that all three genes were involved in the development of female ovules, sterile ovules, and antherophores, but their expression levels, and presumably their roles, differed substantially among these organs. The morphology of transgenic Arabidopsis thaliana (L.) Heynh. flowers suggested that the TM8-like genes had a substantial effect on the emergence and development of short stamens. In addition, the expression patterns of the two transcripts were different and associated with different phenotypes in A. thaliana flowers, suggesting divergent functions in reproductive organ development in G. montanum.
-
Keywords:
- TM8 genes /
- Gnetum montanum /
- Reproductive organ development /
- Arabidopsis thaliana
摘要:TM8基因属于一个古老的Ⅱ型MADS-box基因亚家族,TM8类基因在被子植物中主要参与雌花的发育,但在裸子植物中的功能尚不清楚。本文通过荧光原位杂交FISH(Fluorescence in situ hybridization)和转基因技术分析裸子植物买麻藤(Gnetum montanum Markgr.)3个TM8基因的功能。结果显示,3个基因均参与了雌性胚珠、不育胚珠和花药柄的发育,但其表达水平和功能在器官间有很大差异。将买麻藤TM8基因转入拟南芥(Arabidopsis thaliana (L.) Heynh.),发现其对短雄蕊的萌发和生长有显著影响,其中1个TM8基因的两个转录本呈不同的表达模式,且转基因拟南芥的花呈现不同表型的变化,表明它们在买麻藤生殖器官发育中可能出现了功能分化。
-
-
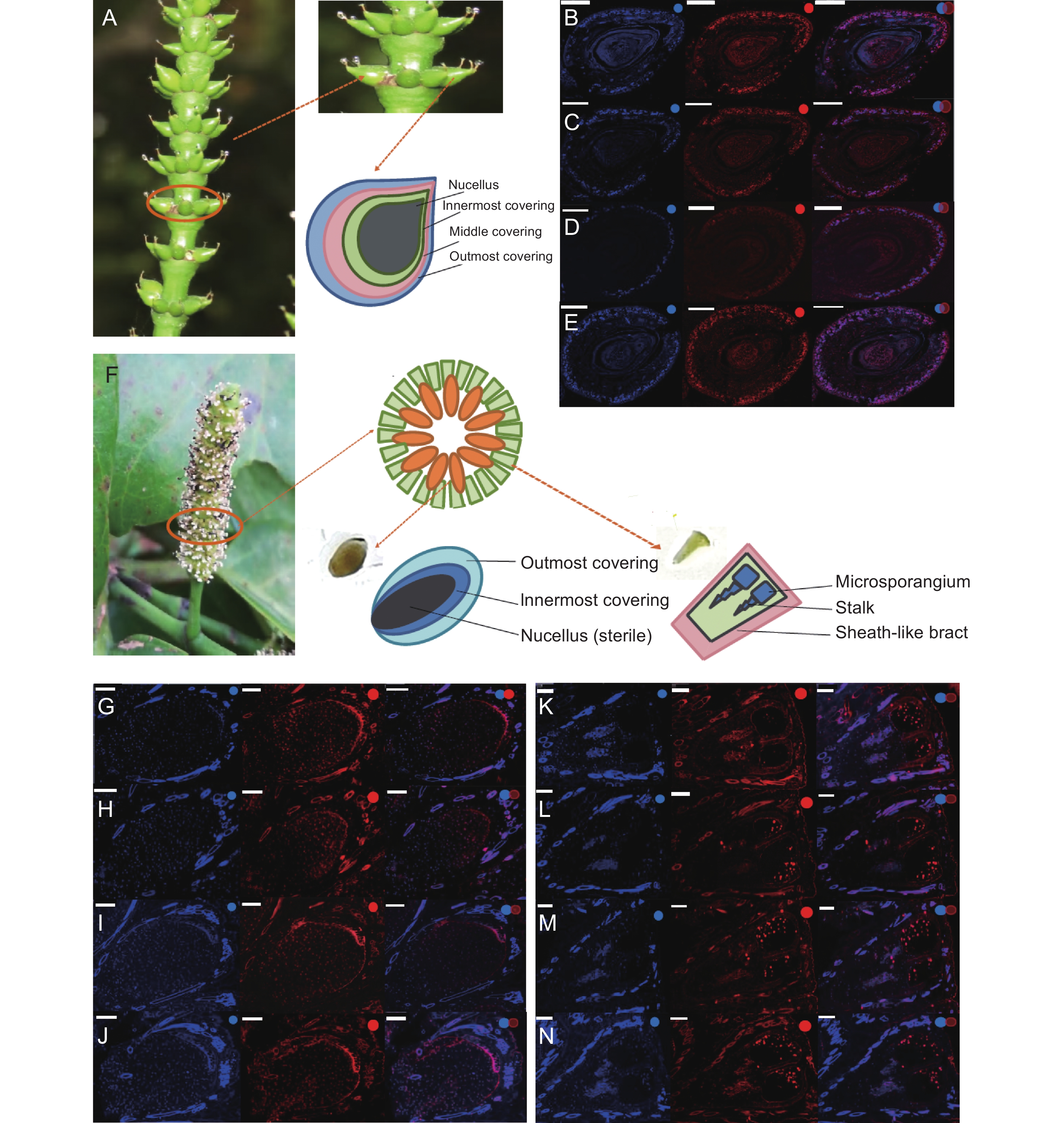
Figure 2. Schematic showing of female and male strobili in Gnetum montanum and results of FISH hybridization
A: Female ovules in a whorl, and anatomical structure of female ovule; B-E: FISH analysis of a female ovule under blue, red, and combined light, white bars represent 500 μm. B: Expression of TnS001008199g01; C: Expression of TnS013912549g01; D: Expression of TnS000980857g03 transcript 1; E: Expression of TnS000980857g03 transcript 2. F: Schematic showing whorls of involucral collars in G. montanum male ovule, sterile ovules and antherophores in a whorl, and anatomical structure of sterile ovule (left) and male reproductive unit (right). G-J: FISH analysis of a sterile ovule under blue, red, and combined light, white bars represent 100 μm. G: Expression of TnS001008199g01; H: Expression of TnS013912549g01; I: Expression of TnS000980857g03 transcript 1; J: Expression of TnS000980857g03 transcript 2. K-N: FISH analysis of antherophores under blue, red, and combined light, white bars represent 100 μm. K: Expression of TnS001008199g01; L: Expression of TnS013912549g01; M: Expression of TnS000980857g03 transcript 1; N: Expression of TnS000980857g03 transcript 2.
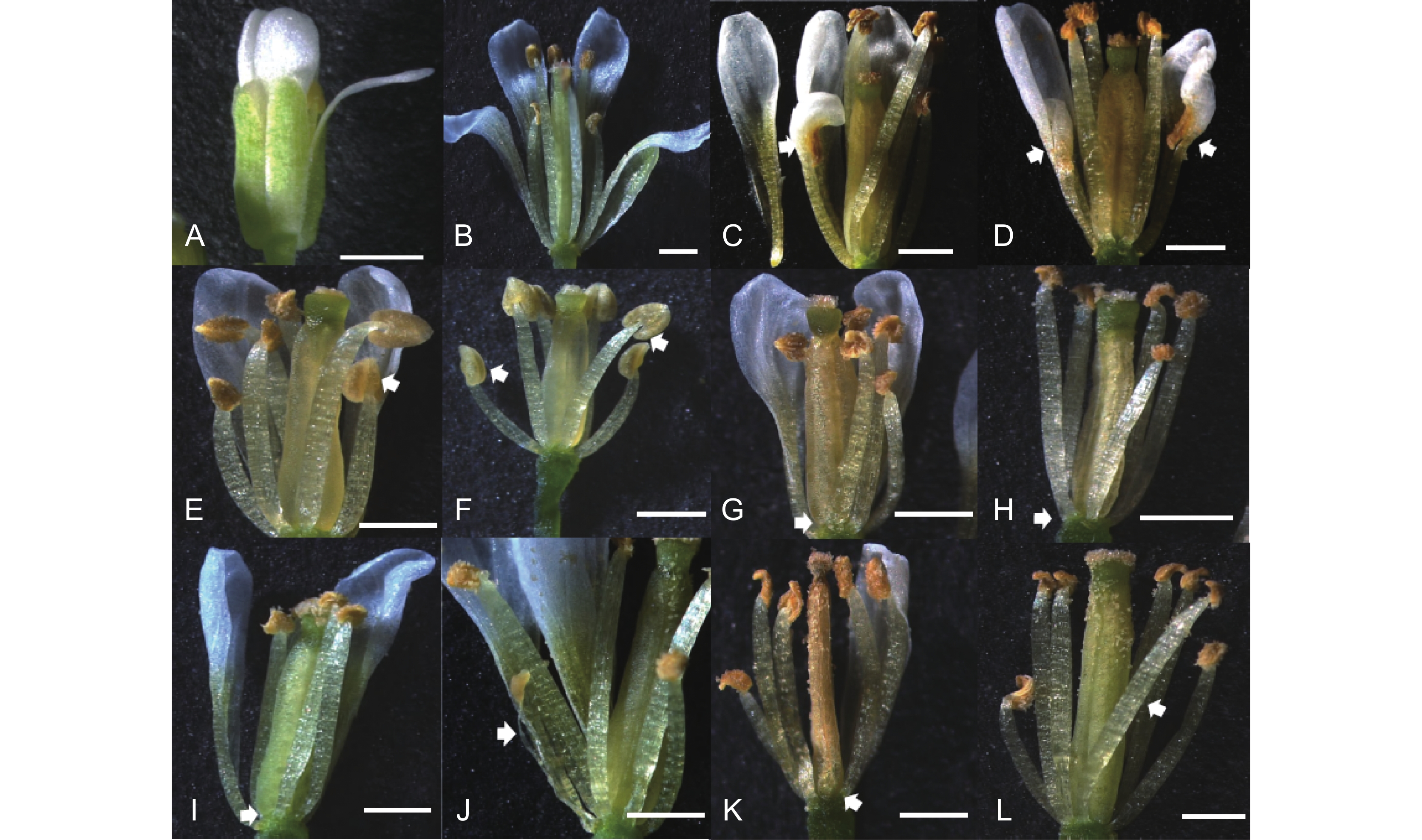
Figure 3. Transgenic experiments of TM8 genes and their transcripts in Arabidopsis thaliana
A, B: Flower of wild-type Arabidopsis thaliana in bloom (a) and unfolded (b); C, D: Floral morphology of TnS001008199g01 transgenic plants from line 1 (c) and line 2 (d), arrows indicate fusion of a petal and stamen; E, F: Floral morphology of TnS013912549g01 transgenic plants from line 1 (e) and line 2 (f), arrows indicate emergence of enlarged stamens; G, H: Floral morphology of TnS000980857g03 transgenic plants from line 1 (g) and line 2 (h), arrows indicate missing short stamens; I, J: Floral morphology of transgenic plants with AS form of TnS000980857g03 from line 1, arrows indicate missing or reduced short stamen; K, L: Floral morphology of transgenic plants with two transcripts of TnS000980857g03 from line 2, arrows indicate missing short stamen or emergence of a fifth long stamen. White bars represent 1 mm.
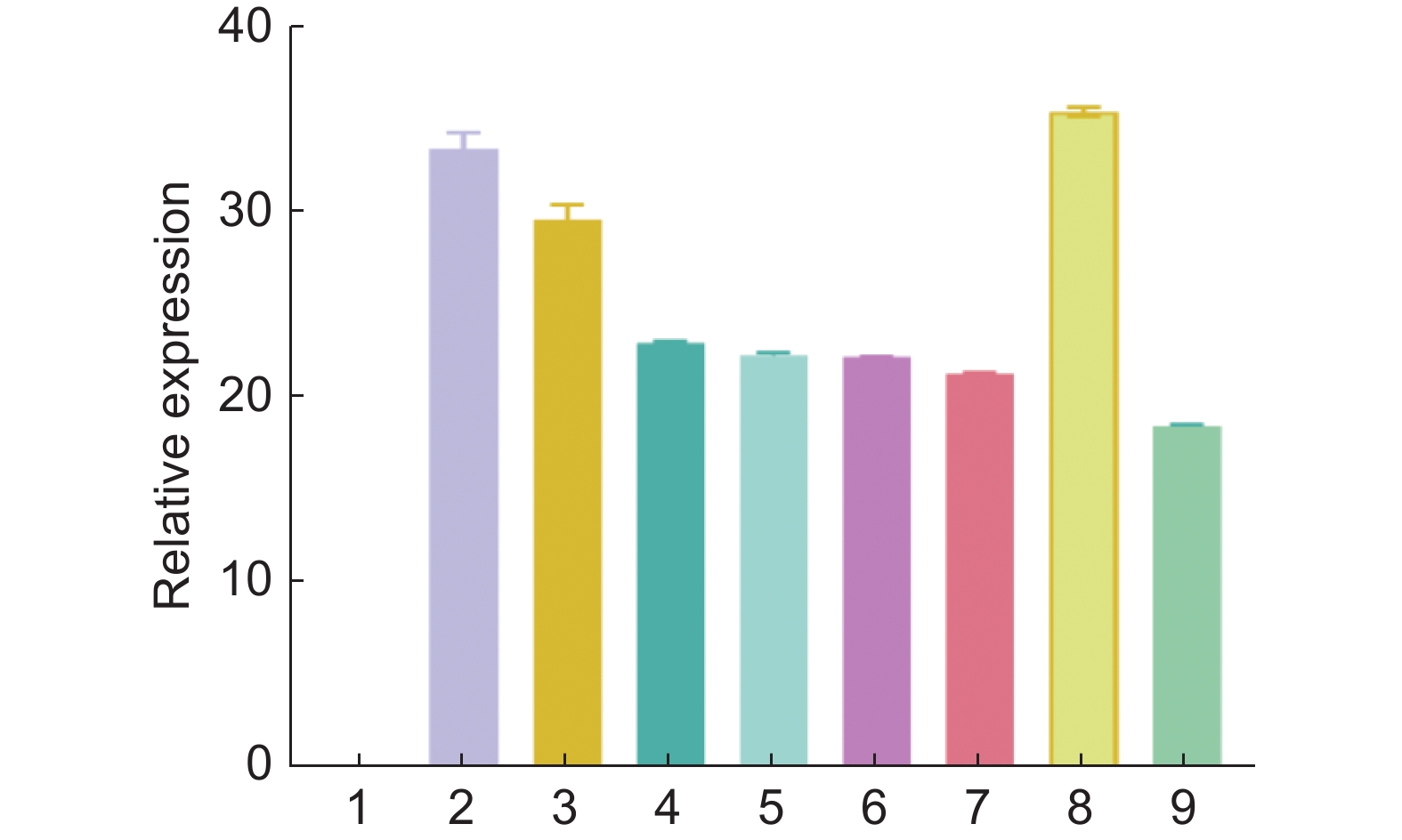
Figure 4. RT-PCR analysis of three TM8-like genes and one AS form
1: Wild type; 2: TnS001008199g01 line 1; 3: TnS001008199g01 line 2; 4: TnS013912549g01 line 1; 5: TnS013912549g01 line 2; 6: TnS000980857g03 line 1; 7: TnS000980857g03 line 2; 8: TnS000980857g03 AS type line 1; 9: TnS000980857g03 AS type line 2.
Table 1 Primers used in the present study
Gene Direction Primer sequence (5'-3') Product length / bp Actin F GCCGACAGAATGAGCAAAGAG 134 R TGCTGGAAGGTACTGAGGGAG TnS001008199g01 F GAAGGATAACGCAAGGCTGAA 255 R TCCTCCTGATTTGCTGCTGTT TnS013912549g01 F GCAGCTCAGCAGGATCGAAAG 291 R TGTAGCCTTTCAATCCGACCA TnS000980857g03
(Transcript 1)F TGTTGCCCTGCTTATTTACTCC 206 R TGCGTACCTCGTCTGCCAAT TnS000980857g03
(Transcript 2)F TCTACCAATTCTCCGATCCAGTG 160 R TCTGCCAATACATGGATCTCCTC -
[1] García-Maroto F,Ortega N,Lozano R,Carmona MJ. Characterization of the potato MADS-box gene STMADS16 and expression analysis in tobacco transgenic plants[J]. Plant Mol Biol,2000,42(3):499−513. doi: 10.1023/A:1006397427894
[2] García-Maroto F,Carmona MJ,Garrido JA,Vilches-Ferrón M,Rodríguez-Ruiz J,López Alonso D. New roles for MADS-box genes in higher plants[J]. Biol Plantarum,2003,46(3):321−330.
[3] Gramzow L,Weilandt L,Theißen G. MADS goes genomic in conifers:towards determining the ancestral set of MADS-box genes in seed plants[J]. Ann Bot,2014,114(7):1407−1429. doi: 10.1093/aob/mcu066
[4] Gramzow L,Theissen G. A hitchhiker's guide to the MADS world of plants[J]. Genome Biol,2010,11(6):214. doi: 10.1186/gb-2010-11-6-214
[5] Melzer R,Wang YQ,Theißen G. The naked and the dead:the ABCs of gymnosperm reproduction and the origin of the angiosperm flower[J]. Semin Cell Dev Biol,2010,21(1):118−128. doi: 10.1016/j.semcdb.2009.11.015
[6] Gramzow L,Theißen G. Phylogenomics reveals surprising sets of essential and dispensable clades of MIKCc-group MADS-box genes in flowering plants[J]. J Exp Zool B Mol Dev Evol,2015,324(4):353−362. doi: 10.1002/jez.b.22598
[7] Becker A,Theißen G. The major clades of MADS-box genes and their role in the development and evolution of flowering plants[J]. Mol Phylogen Evol,2003,29(3):464−489. doi: 10.1016/S1055-7903(03)00207-0
[8] Heijmans K,Morel P,Vandenbussche M. MADS-box genes and floral development:the dark side[J]. J Exp Bot,2012,63(15):5397−5404. doi: 10.1093/jxb/ers233
[9] Pnueli L,Abu-Abeid M,Zamir D,Nacken W,Schwarz-Sommer Z,Lifschitz E. The MADS box gene family in tomato:temporal expression during floral development,conserved secondary structures and homology with homeotic genes from Antirrhinum and Arabidopsis[J]. Plant J,1991,1(2):255−266. doi: 10.1111/j.1365-313X.1991.00255.x
[10] Ando S,Sato Y,Kamachi S,Sakai S. Isolation of a MADS-box gene (ERAF17) and correlation of its expression with the induction of formation of female flowers by ethylene in cucumber plants (Cucumis sativus L. )[J]. Planta,2001,213(6):943−952. doi: 10.1007/s004250100571
[11] Daminato M,Masiero S,Resentini F,Lovisetto A,Casadoro G. Characterization of TM8,a MADS-box gene expressed in tomato flowers[J]. BMC Plant Biol,2014,14:319. doi: 10.1186/s12870-014-0319-y
[12] Lovisetto A,Guzzo F,Tadiello A,Toffali K,Favretto A,Casadoro G. Molecular analyses of MADS-box genes trace back to gymnosperms the invention of fleshy fruits[J]. Mol Biol Evol,2012,29(1):409−419. doi: 10.1093/molbev/msr244
[13] Hou C,Humphreys AM,Thureborn O,Rydin C. New insights into the evolutionary history of Gnetum (Gnetales)[J]. Taxon,2015,64(2):239−253. doi: 10.12705/642.12
[14] Hou C,Wikström N,Strijk JS,Rydin C. Resolving phylogenetic relationships and species delimitations in closely related gymnosperms using high-throughput NGS,sanger sequencing and morphology[J]. Plant Syst Evol,2016,302(9):1345−1365. doi: 10.1007/s00606-016-1335-1
[15] Hou C,Li LF,Liu ZM,Su YJ,Wan T. Diversity and expression patterns of MADS-box genes in Gnetum luofuense: implications for functional diversity and evolution[J]. Trop Plant Biol,2020,13(1):36−49. doi: 10.1007/s12042-019-09247-x
[16] Deng N,Hou C,He BX,Ma FF,Song QG,et al. A full-length transcriptome and gene expression analysis reveal genes and molecular elements expressed during seed development in Gnetum luofuense[J]. BMC Plant Biol,2020,20(1):531. doi: 10.1186/s12870-020-02729-1
[17] Barbazuk WB,Fu Y,McGinnis KM. Genome-wide analyses of alternative splicing in plants:opportunities and challenges[J]. Genome Res,2008,18(9):1381−1392. doi: 10.1101/gr.053678.106
[18] Kalsotra A,Cooper TA. Functional consequences of developmentally regulated alternative splicing[J]. Nat Rev Genet,2011,12(10):715−729. doi: 10.1038/nrg3052
[19] Wang BB,Brendel V. Genomewide comparative analysis of alternative splicing in plants[J]. Proc Natl Acad Sci USA,2006,103(18):7175−7180. doi: 10.1073/pnas.0602039103
[20] Ye JB,Cheng SY,Zhou X,Chen ZX,Kim SU,et al. A global survey of full-length transcriptome of Ginkgo biloba reveals transcript variants involved in flavonoid biosynthesis[J]. Ind Crop Prod,2019,139:111547. doi: 10.1016/j.indcrop.2019.111547
[21] Hou C,Deng N,Su YJ. PacBio long-read sequencing reveals the transcriptomic complexity and Aux/IAA gene evolution in Gnetum (Gnetales)[J]. Forests,2019,10(11):1043. doi: 10.3390/f10111043
[22] Hou C,Lian HM,Cai YL,Wang YL,Liang DC,He BX. Comparative analyses of full-length transcriptomes reveal Gnetum luofuense stem developmental dynamics[J]. Front Genet,2021,12:615284. doi: 10.3389/fgene.2021.615284
[23] Lalitha S. Primer premier 5[J]. Biotech Software Internet Rep,2000,1(6):270−272. doi: 10.1089/152791600459894
[24] Livak KJ,Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔ CT method[J]. Methods,2001,25(4):402−408. doi: 10.1006/meth.2001.1262
[25] Berridge EM. On some points of resemblance between gnetalean and Bennettitean seeds[J]. New Phytol,1911,10(4):140−144. doi: 10.1111/j.1469-8137.1911.tb04959.x
[26] Pitto L,Giorgetti L,Turrini A,Evangelista M,Luccarini G,et al. Floral genes expressed in tomato hypocotyl explants in liquid culture[J]. Protoplasma,2001,218(3-4):168−179. doi: 10.1007/BF01306606
[27] Shindo S,Ito M,Ueda K,Kato M,Hasebe M. Characterization of MADS genes in the gymnosperm Gnetum parvifolium and its implication on the evolution of reproductive organs in seed plants[J]. Evol Dev,1999,1(3):180−190. doi: 10.1046/j.1525-142x.1999.99024.x
[28] Endress PK. Structure and function of female and bisexual organ complexes in Gnetales[J]. Int J Plant Sci,1996,157(S6):S113−S125. doi: 10.1086/297407
[29] Winter KU,Becker A,Münster T,Kim JT,Saedler H,Theissen G. MADS-box genes reveal that gnetophytes are more closely related to conifers than to flowering plants[J]. Proc Nat Acad Sci USA,1999,96(13):7342−7347. doi: 10.1073/pnas.96.13.7342
[30] Becker A,Saedler H,Theissen G. Distinct MADS-box gene expression patterns in the reproductive cones of the gymnosperm Gnetum gnemon[J]. Dev Genes Evol,2003,213(11):567−572. doi: 10.1007/s00427-003-0358-0


 下载:
下载: